Landmark Parkinson's research and other major studies contain obvious data errors that went undetected for years, raising questions about scientific integrity and oversight.
A landmark Parkinson's disease study that claimed the disease originates in the gut rather than the brain contains obvious copy-paste errors in its underlying data, according to new analysis. The paper, which has received over 3000 citations and major media coverage, has had its dataset publicly available on Dryad for more than 8 years without anyone noticing the blatant errors.
The errors were discovered by a software tool designed to detect copy-paste patterns in scientific datasets. The tool was inspired by two high-profile cases of data fabrication that made headlines in recent years - one involving Nobel laureate Thomas Südhof's lab and another by spider ecologist Jonathan Pruitt. Both cases had publicly available datasets with entire blocks of copy-pasted data that seemed trivial to detect.
"I was curious what I could dig up by creating a program that would correctly flag those cases, and then unleash it on all datasets available in open-access repositories," explains the researcher behind the tool.
Case 1: The Parkinson's paper
The study in question, published in Cell in 2016, claimed that clearing out the microbiome of mice genetically predisposed to Parkinson's symptoms made all their symptoms disappear. The senior author Sarkis Mazmanian summarized the findings on the Rich Roll YouTube channel, stating that mice with a normal gut microbiome developed symptoms while those without a microbiome remained healthy.
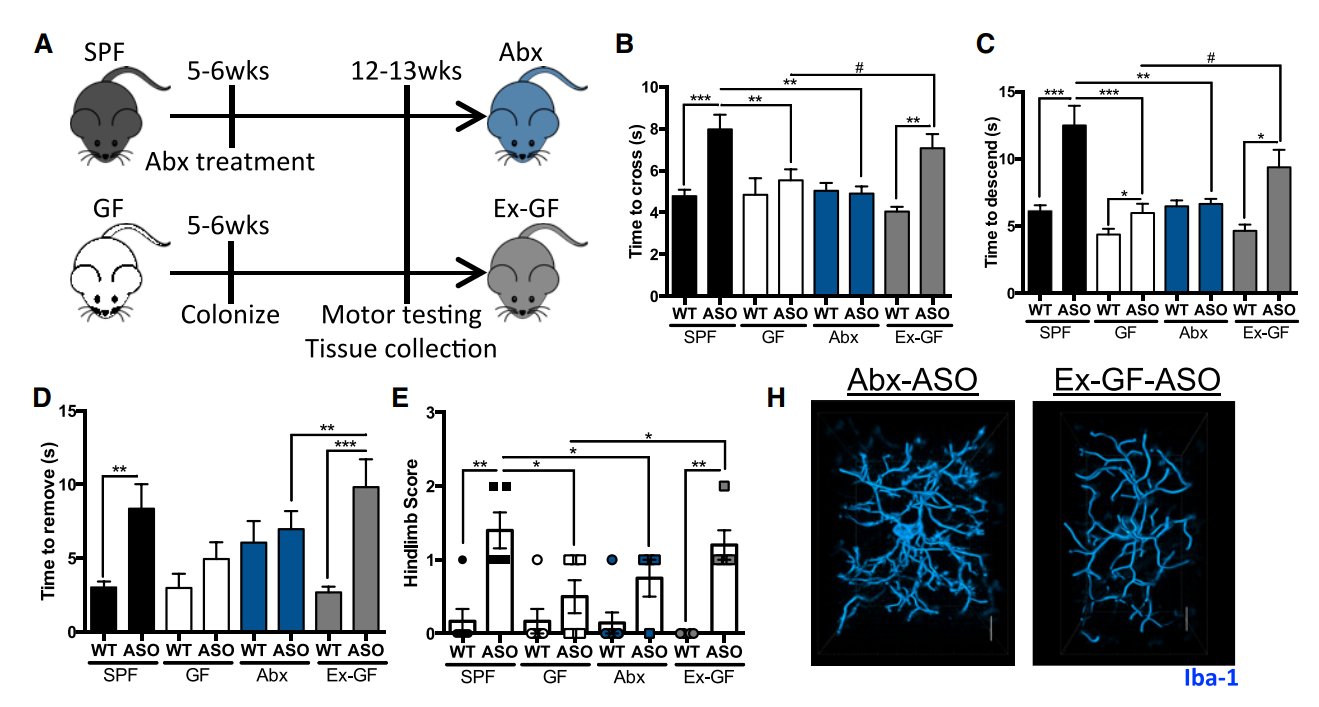
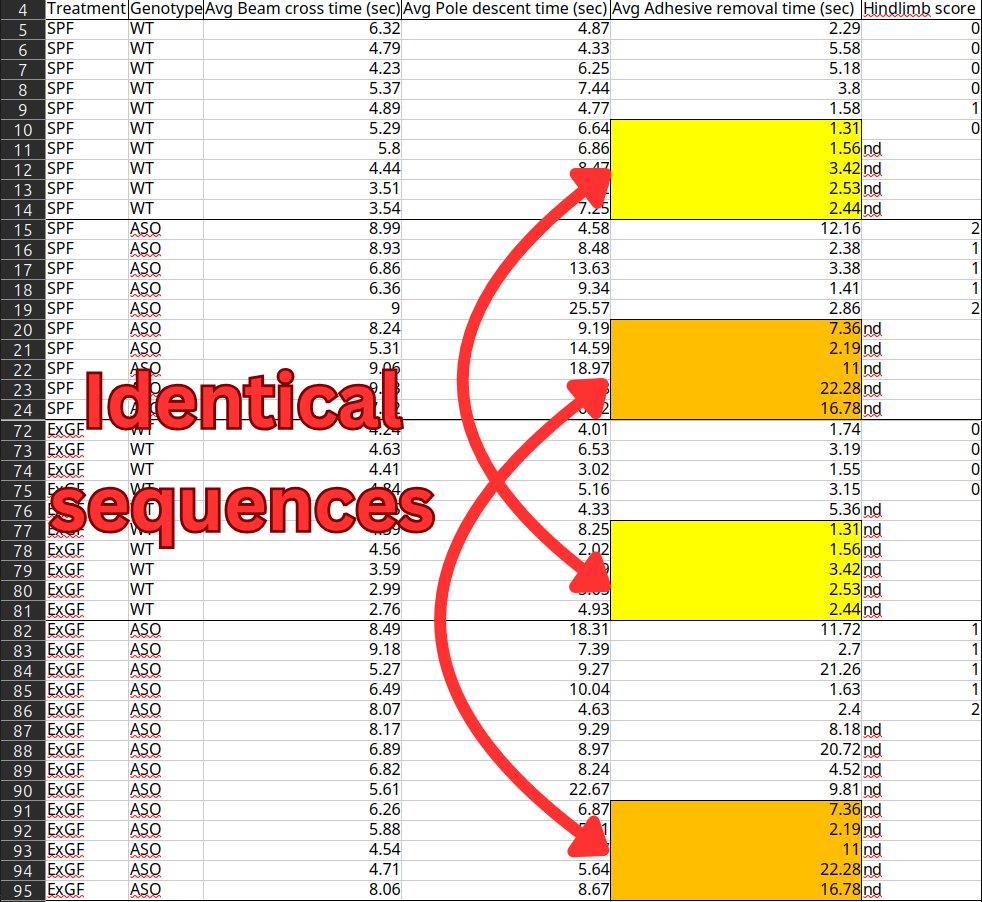
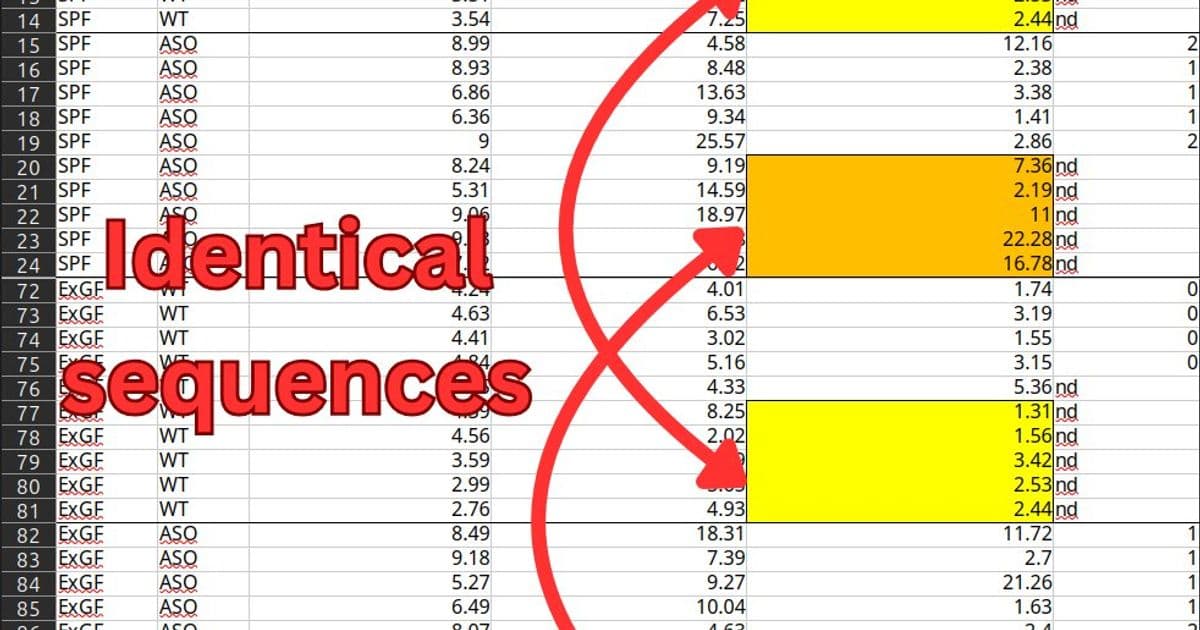
However, the dataset contains two sets of 5 identical sequential numbers in the "Adhesive removal times" column, shared between different groups of mice. There's also a pair of sequences of 3 identical numbers within the "Pole-descent time" data for germ-free wild-type mice.
"The duplicated rows make up 50% of the SPF samples and 42% of the ExGF samples," the analysis notes. "While the motor function values are not the only data the authors rely on for their conclusions, they're necessary for showing that the gut bacteria actually caused Parkinson's-like symptoms."
The issue was reported in January, but the authors have not responded.
Case 2: The ostrich-snake mixup
A 2022 study in PLOS Genetics investigated how different animals evolved resistance to cardiotonic steroids - toxins that prey species produce to protect themselves against predators like birds and snakes.
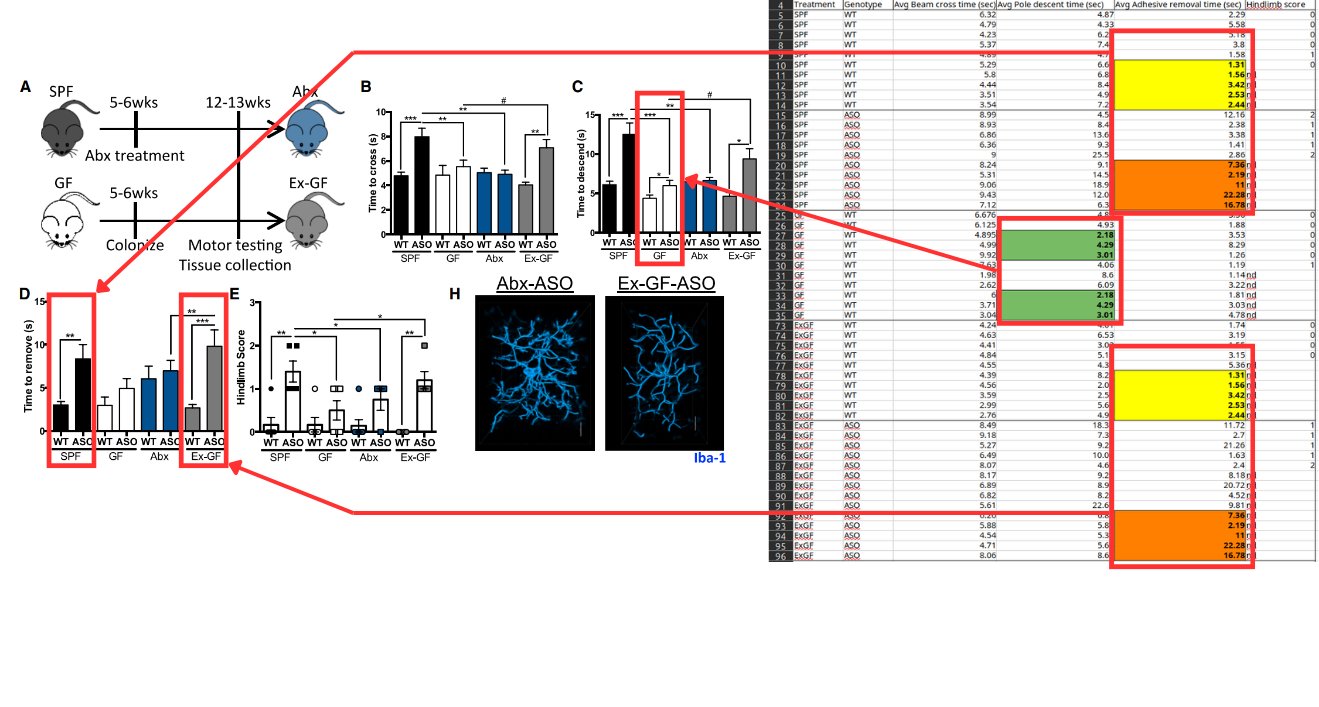
The dataset contains precise duplicates between ostrich/sandgrouse data and snake data, plus near-duplicates that differ by one or two digits but always end with the same digit. For example, 0.538 becomes 0.518, with six such pairs out of eight total non-duplicate pairs.
The lead author Shabnam Mohammadi initially suggested these could be measurement variations from multiple reads of the same plate. However, the pattern where values end with the same digit is statistically unlikely to occur by chance.
"The alternative explanation is that the authors either accidentally or deliberately copy-pasted the ostrich/sandgrouse data on top of the snake data," the analysis suggests. "Then they might have manually tampered with the data to make it better suit their hypothesis or make it look less suspicious."
Case 3: Scrambled fish sizes
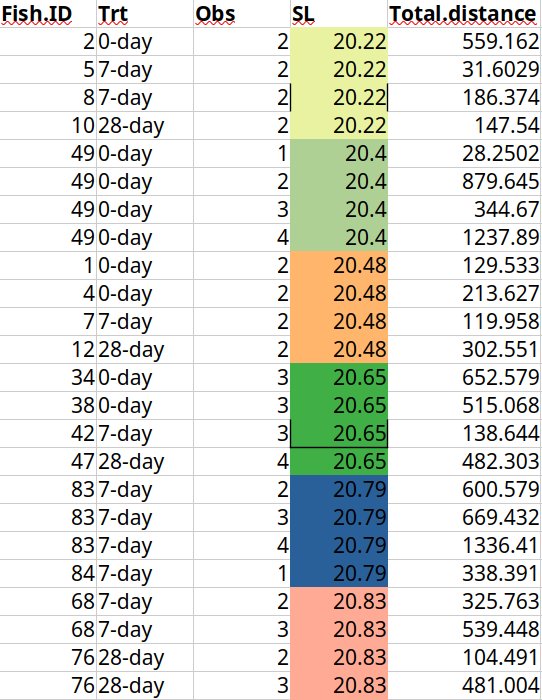
A 2017 Nature Communications study on fish personalities used genetically identical fish to examine behavioral differences. The dataset shows three different individual fish all having the exact same sequential size measurements.
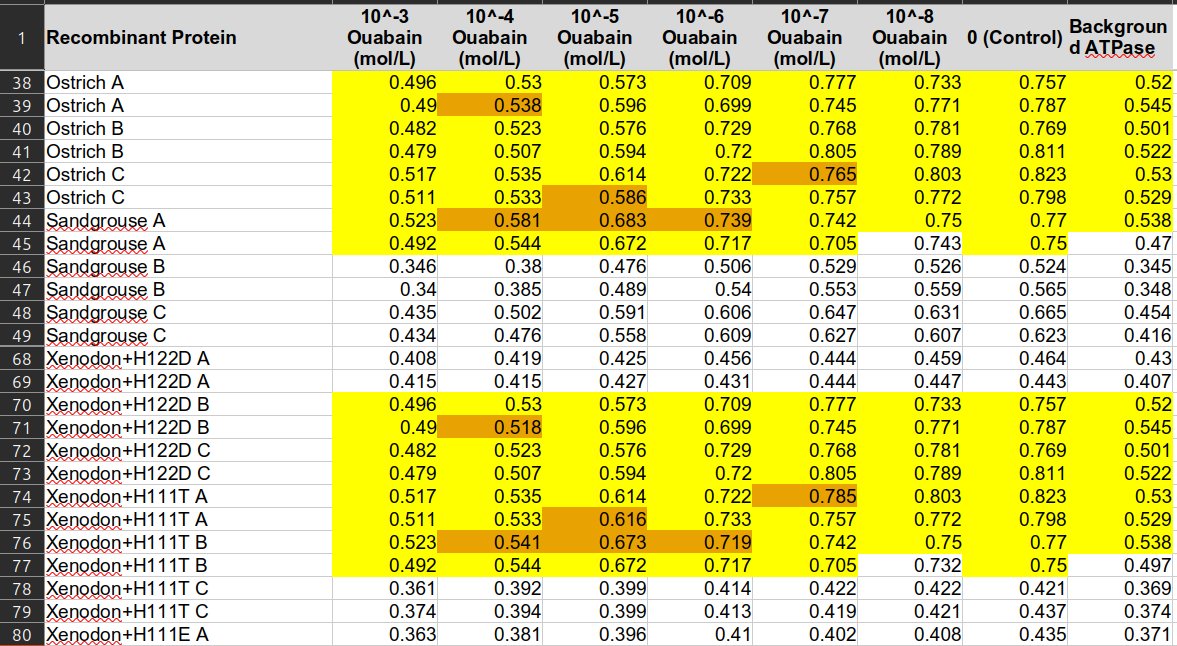
Further investigation revealed that every unique fish length value reoccurs exactly four times - matching the number of observations done per individual fish. The authors admitted they accidentally misaligned ID values when joining separate data files, causing all body size values to be shifted and assigned to wrong rows.
The scale of the problem
The software detected 18 serious cases in the first 600 datasets scanned, with 15 posted to PubPeer. Based on this limited sample, around 3% of papers contain these types of errors. However, the true error rate is likely much higher.
"There are myriad other ways of accidentally screwing up your data that this software could never detect," the researcher notes. "And if you want to commit fraud, there are plenty of less lazy ways to do it than to copy-paste values from a different part of the same Excel sheet."
The lack of detection raises questions about scientific oversight. "It might come as a surprise to some that nobody else has cared to check for these errors," the analysis states. "Isn't there some peer review thing that is supposed to prevent stuff like this from making it into the scientific record?"
The conclusion is stark: "There just isn't anybody whose job it is to actively look for it. Journals, universities and funding organizations won't hire anyone to do it because they generally care a lot more about rankings and metrics."
What's next
The project has received a $50,000 grant from Astral Codex Ten, allowing full-time work on scanning through the remaining ~24,000 datasets with Excel files available on Dryad. At the current 3% error rate, that could reveal around 700 more cases.
"It will be interesting to see what other treasures we can dig up there!" the researcher concludes. The findings highlight a critical gap in scientific quality control and raise serious questions about the reliability of published research when basic data errors can go undetected for years.
Comments
Please log in or register to join the discussion