MIT researchers have quantified chromatin movement across unprecedented timescales, revealing two distinct dynamic classes that govern how genes interact with regulatory elements, with significant implications for understanding gene expression and DNA repair mechanisms.
For decades, biologists have known that gene expression depends on interactions between genes and regulatory elements along the genome. However, the precise mechanics of how these interactions occur within the crowded environment of the cell nucleus have remained poorly understood. In a groundbreaking study published in Nature Structural and Molecular Biology, MIT researchers have successfully measured chromatin movement across an unprecedented range of timescales, providing the first rigorous quantification of chromatin dynamics and revealing how these movements facilitate essential biological processes.
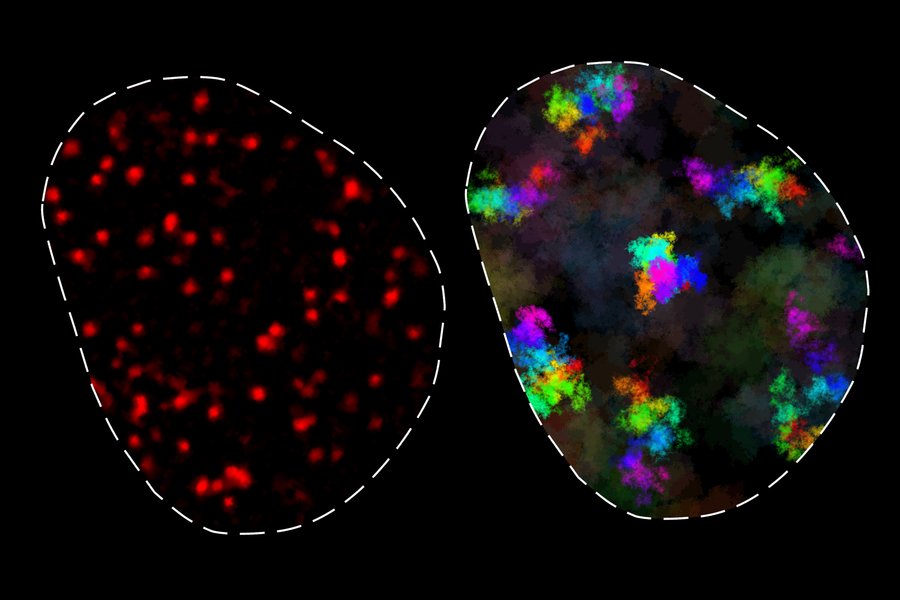
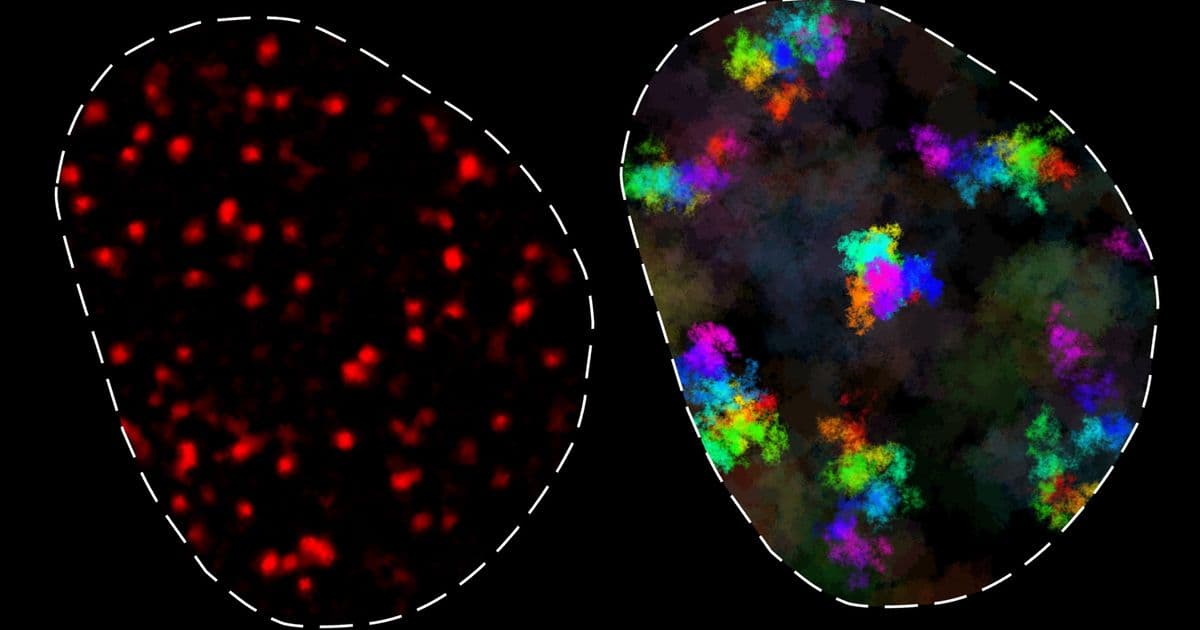
The research team, led by Associate Professor Anders Sejr Hansen, discovered that chromatin exists in two distinct dynamic classes. In one state, chromatin moves in a constrained manner, primarily contacting only neighboring genomic regions. In the other state, chromatin moves more freely, enabling contact with distant genomic elements, but only over longer timescales. These findings fundamentally advance our understanding of how genes locate and interact with their regulatory elements—a process critical for proper cellular function.
"Because we were able to look at chromatin dynamics for the first time at these very fast timescales, and also for the first time across the full dynamic range, we were able to observe chromatin motion over a range that just wasn't possible before," explains Hansen, senior author of the study.
Technical Approach: Breaking the Time Scale Barrier
Previous studies of chromatin dynamics yielded conflicting results, primarily due to limitations in tracking technology. Conventional microscopy techniques could only observe chromatin movement over limited time ranges, making it difficult to obtain statistically robust measurements. The MIT team overcame this limitation by employing MINFLUX (minimal fluorescence fluxes), a super-resolution light microscopy technique developed by Nobel laureate Stefan Hell of the Max Planck Institute.
"MINFLUX allowed us to get around the limitations of conventional microscopy, letting us measure chromatin movement faster and for a longer period of time than ever before," says Domenic Narducci, one of the lead authors. "To our knowledge, it's the first time this technique has been used this way."
By combining MINFLUX with traditional imaging techniques, the researchers successfully tracked chromatin movement across seven orders of magnitude in time—from hundreds of microseconds to several hours. This comprehensive temporal coverage revealed patterns that had previously been invisible to researchers.
The study was conducted across multiple mouse and human cell types, allowing the researchers to identify not only the fundamental dynamics of chromatin movement but also how these dynamics vary between different cellular environments.
The Physics of Chromatin Movement
Contrary to textbook depictions of chromatin as a static structure, the researchers found that chromatin is in constant motion. This movement is constrained by the fundamental physics of DNA as a polymer. When a particular location on the genome (locus) moves, it experiences a pulling force from the DNA segments on either side.
"Chromosomes are polymers. They're held together by many nucleotides of DNA. Being part of DNA is a little bit like running while holding hands with other people. If a hundred people are holding hands and you, in the middle of the chain, try to run in one direction, you'll get pulled back," Hansen explains.
This behavior, known as subdiffusive movement, means that chromatin doesn't move freely but is instead constrained by its polymer nature. The MIT researchers found that this constraining effect is stronger than previously thought, with any given locus typically moving only within about 200 nanometers over short and intermediate timescales (up to 200 seconds).
"One of the main takeaways is that you have this region of influence where a genomic locus has access to other genomic loci, and this is roughly a couple hundred nanometers large," says Simon Grosse-Holz, another lead author. "If loci are much closer together than a couple hundred nanometers, they're effectively in contact all the time. You get a cutoff at a couple hundred nanometers where everything within that region around a given locus can see that locus, and everything outside cannot."
Implications for Gene Regulation and DNA Repair
The findings have significant implications for understanding how genes interact with regulatory elements. The researchers discovered that for genes and regulatory elements within approximately 100,000 base pairs of each other, no special mechanisms are needed to facilitate their interaction—they will naturally find each other through normal chromatin movement.
"If they are closer than 100,000 bases, and most regulatory elements are, then those elements are going to find their target gene within a few milliseconds or a few minutes," explains Matteo Mazzocca, the third lead author. "These are timescales that are completely consistent with transcription."
This constant proximity within the 200-nanometer "region of influence" is particularly important for DNA repair. When DNA breaks occur, the two ends of the DNA remain in close proximity, facilitating their eventual reunion and repair.
The second class of chromatin dynamics—characterized by wider movement but only over longer timescales (minutes to hours)—was observed in some cell types but not others. This variation between cell types was unexpected and suggests that chromatin dynamics may be more specialized than previously believed.
"It would be reasonable to assume that the behavior would be more or less the same in all cell types, but that's not at all what we found," Hansen notes. "It's very different in different cell types, with no obvious way of categorizing things."
Challenging Existing Models
The strength of the subdiffusive pull observed in this study cannot be explained by existing models of chromatin dynamics—the Rouse model and the fractal globule model. This suggests that these models may need to incorporate additional factors, such as the interactions between chromatin and the crowded nucleoplasm environment in which it exists.
"These findings are significant for two key reasons," comments Luca Giorgetti, a group leader at the Friedrich Miescher Institute for Biomedical Research in Switzerland, who was not involved in the study. "First, they rigorously confirm longstanding but anecdotal observations that chromatin motion is strongly subdiffusive. Second, they demonstrate that this behavior is consistent across multiple cell types and persists across all measured timescales."
Future Directions
The research opens numerous avenues for future investigation. Understanding how chromatin dynamics vary between cell types could provide insights into cellular specialization and dysfunction in diseases like cancer. Additionally, the MINFLUX technique used in this study could be applied to study other dynamic processes within the cell nucleus.
The researchers are particularly interested in exploring how chromatin dynamics change during different stages of the cell cycle and in response to cellular stress or damage. They also hope to investigate how specific proteins and modifications might influence chromatin movement and, consequently, gene expression.
The study was funded by the National Institutes of Health, a National Science Foundation CAREER Award, a Pew-Stewart Scholar for Cancer Research Award, and the Bridge Project, a partnership between the Koch Institute for Integrative Cancer Research at MIT and the Dana-Farber/Harvard Cancer Center.
For researchers interested in exploring this work further, the original paper titled "Integrated MINFLUX tracking reveals two distinct chromatin dynamics classes across cell types" is available in Nature Structural and Molecular Biology. The research team has also made their methodologies and data analysis approaches available through their laboratory website at MIT.
This research represents a significant step forward in understanding the physical principles that govern gene expression and provides a foundation for future studies into how disruptions in chromatin dynamics might contribute to disease and developmental disorders.
Comments
Please log in or register to join the discussion