MIT scientists have discovered that the physical arrangement of genes along DNA strands creates structural changes that dramatically affect neighboring gene expression, opening new possibilities for synthetic biology applications.
MIT Research Reveals Gene 'Syntax' Controls DNA Structure and Gene Expression
When a gene is activated within a cell, it sends physical ripples through the DNA strand that can either enhance or suppress neighboring genes, according to new research from MIT. This discovery reveals that the relative positioning of genes—what the researchers call "gene syntax"—plays a crucial role in determining how genes are expressed, offering new tools for designing synthetic biological systems.
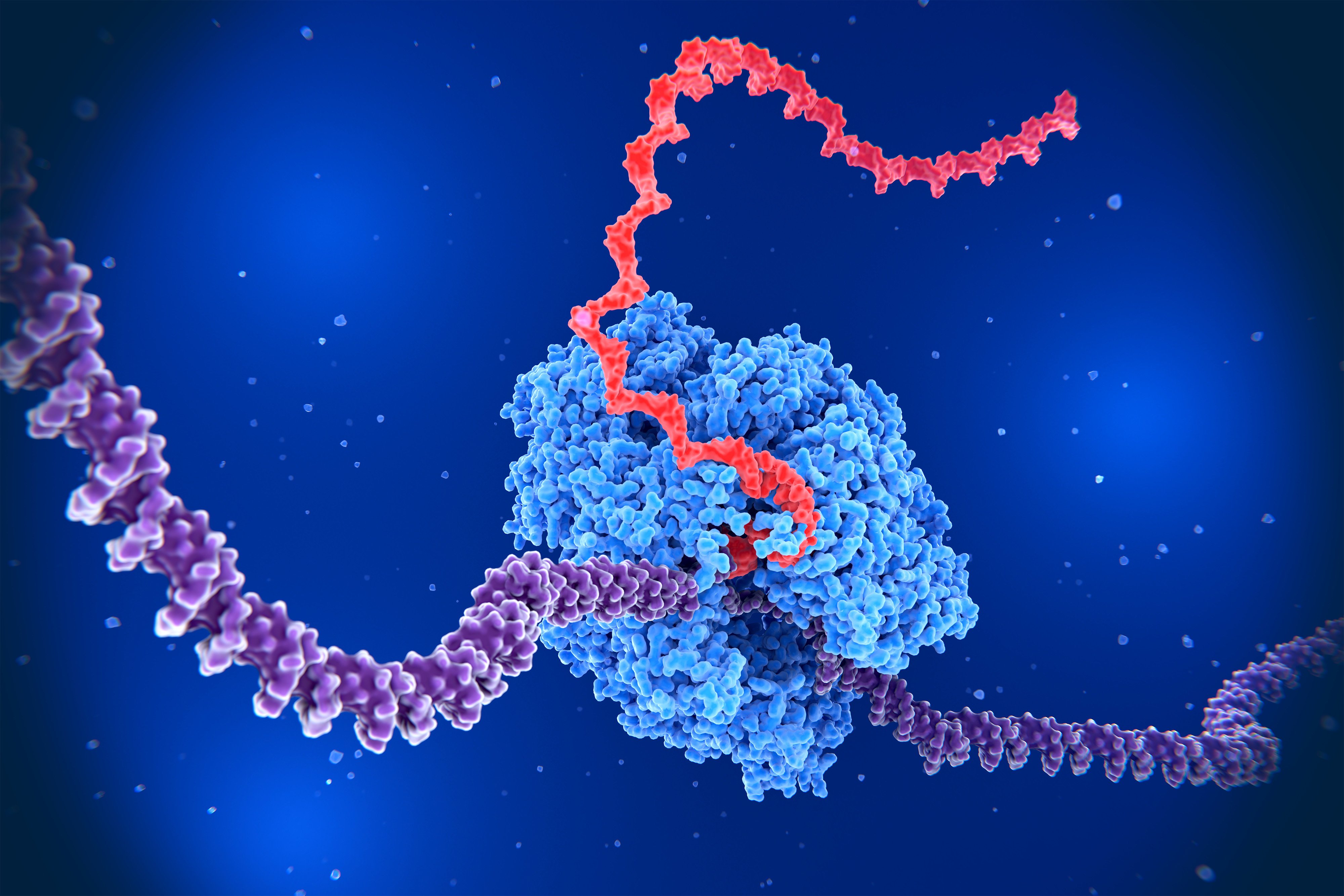
Physical Changes in DNA Structure
The MIT team, led by Assistant Professor Katie Galloway, found that when a gene is transcribed into messenger RNA, the double-stranded DNA helix must unwind to allow RNA polymerase enzyme access to the genetic code. This unwinding process creates distinct physical changes along the DNA strand:
- Upstream regions: DNA becomes more relaxed, making it easier for RNA polymerase to bind and initiate transcription
- Downstream regions: DNA becomes tightly wound into structures called plectonemes (similar to tangled telephone cords), making it harder for RNA polymerase to access the DNA
"Normally, we think about gene circuits and pieces of DNA as these lines that we draw, but they're polymers that have physical characteristics," Galloway explains. "The thing that we were trying to solve in this paper was: When you put two genes on the same piece of DNA, how does their physical interaction become coupled?"
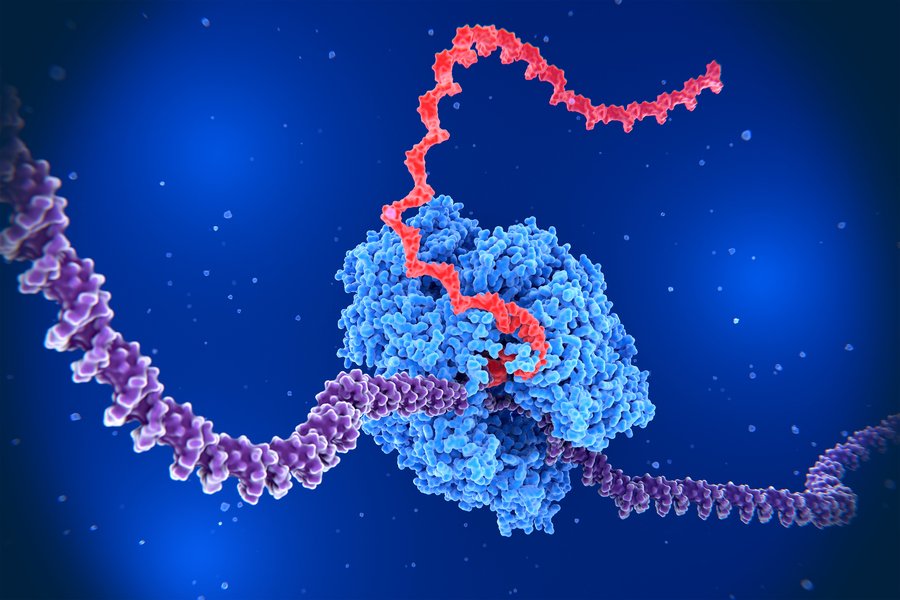
Gene Syntax and Expression Patterns
The researchers studied three different gene arrangements or "syntaxes":
- Tandem arrangement: Genes positioned sequentially in the same direction
- Divergent arrangement: Neighboring genes transcribed in opposite directions (away from each other)
- Convergent arrangement: Genes transcribed toward each other
Through both computational modeling and experimental validation in human cells and induced pluripotent stem cells, the team confirmed that these arrangements produce dramatically different expression patterns:
- Divergent syntax: Both genes are expressed at high levels, potentially amplifying output
- Tandem syntax: The upstream gene suppresses expression of the downstream gene
- Convergent syntax: Not explicitly detailed in the findings
These effects were substantial, producing as much as a 25-fold increase or decrease in gene expression, and were observable at distances of up to 2,000 base pairs between genes.
Experimental Validation
To validate their computational predictions, the researchers engineered synthetic gene circuits with different syntaxes into human cell lines. They used a high-resolution genome mapping technique called Region Capture Micro-C to analyze DNA structural changes during transcription.
"In divergent circuits, expression of both genes was amplified. In tandem circuits, turning on the upstream gene suppressed the expression of the downstream gene," the researchers reported.
The team also developed a new system called STRAIGHT-IN Dual in collaboration with Leiden University Medical Center researchers, which allows efficient insertion of two genes into the same DNA strand at both alleles. This system is described in a companion paper published in Nature Biomedical Engineering.
Applications in Synthetic Biology
These findings have significant implications for synthetic biology and genetic engineering:
- Enhanced circuit design: Researchers can now consider both biochemical and biophysical factors when designing synthetic gene circuits
- Optimized protein production: The divergent syntax could be used to maximize production of therapeutic proteins like antibodies
As a demonstration, the researchers built synthetic circuits containing genes for two segments of a novel antibody used to treat yellow fever. As predicted, the divergent syntax produced larger quantities of the therapeutic antibody.
"This is really exciting because we can coordinate gene expression in ways that just weren't possible before," Galloway says. "Syntax will be really useful for dynamic circuits. Now we have the ability to select not only the biochemistry of circuits, but also the physical design to support dynamics."
Future Directions
The research team is already applying these principles to their previous work on gene delivery systems and cell reprogramming techniques. This gene syntax approach could enable:
- Toggle switches that alternate between two gene states
- Oscillators that produce periodic gene expression patterns
- Pulse generators for timed release of therapeutic proteins
"If you want coordinated expression, a divergent circuit is great. If you want something that's either/or, you can imagine using a convergent or tandem circuit, so when one turns on, the other turns off, and you can alternate pulses," Galloway explains. "Now that we understand the syntax, I think this will pave the way for us to program dynamic behaviors."
The research was supported by the National Institutes of Health, National Science Foundation, and other funding agencies. The findings were published in the journal Science, with a related paper on the STRAIGHT-IN Dual system appearing in Nature Biomedical Engineering.
For more information on the Galloway Lab's research, visit their official page. The research paper "Gene syntax defines supercoiling-mediated transcriptional feedback" is available in the current issue of Science.
Comments
Please log in or register to join the discussion