MIT researchers have identified specific bacterial signals that neurons detect, providing fundamental insights into how the gut microbiome influences brain function through neural interactions.
In the intricate relationship between animals and their bacterial companions, a groundbreaking study from MIT's Picower Institute for Learning and Memory has revealed the precise mechanisms by which neurons detect and respond to bacteria in the gut. The research, published in Current Biology and led by Picower Fellow Cassi Estrem in Associate Professor Steven Flavell's lab, identifies specific chemical compounds that trigger key neural responses in the nematode Caenorhabditis elegans, offering fundamental insights that may extend to more complex organisms, including humans.
The Human Microbiome Connection
The human gut microbiome has been increasingly associated with various neurological conditions, including depression and Parkinson's disease. However, understanding the actual mechanisms that enable bacterial communities to influence brain function has remained challenging. Flavell, an investigator of the Howard Hughes Medical Institute and faculty member of MIT's Department of Brain and Cognitive Sciences, emphasizes the significance of this relationship: "In our bodies, our own cells are outnumbered by the bacterial cells living in and on us. There's an increasing recognition that this has a profound impact on human health. It's been clear that there are links for some time. Our study aimed to identify the hard mechanisms of how a host nervous system is affected by bacteria in the alimentary canal."
By studying C. elegans—a "bacterial specialist" that evolved to eat bacteria as its primary diet while avoiding pathogenic bacteria—the researchers gained insights into a nervous system particularly attuned to distinguishing between beneficial and harmful bacteria. Understanding these fundamental mechanisms could lead to improved therapeutic interventions targeting the microbiome-brain axis.
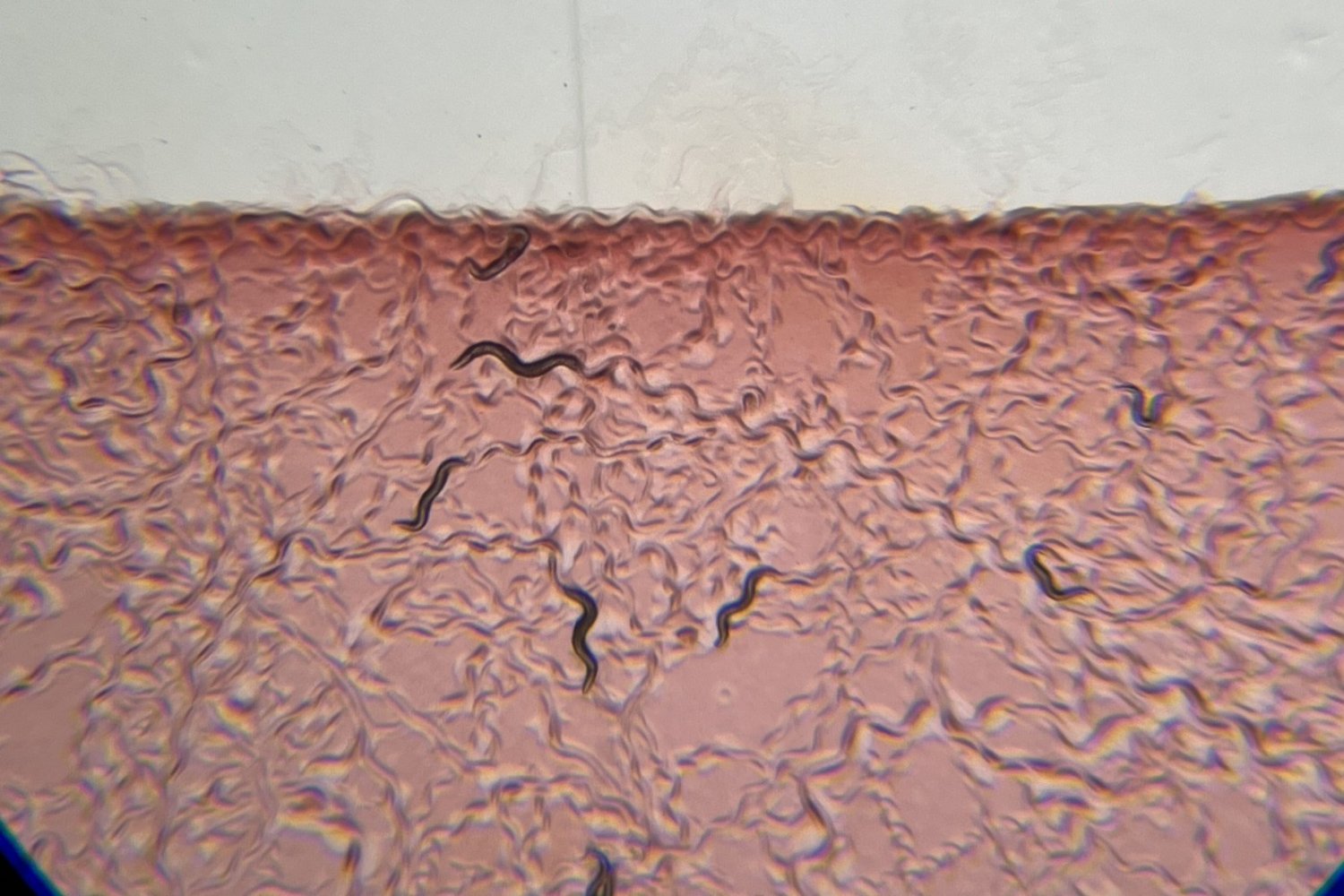

The NSM Neuron and Its Detection Mechanism
In 2019, Flavell's lab discovered that the NSM neuron, which projects into the worm's alimentary canal, employs two "acid sensing ion channels" (ASICs) to detect ingested bacteria. These ion channels are analogous to those found in human neurons. When NSM detects beneficial bacteria, it releases serotonin, causing the worm to increase its feeding rate and slow its movement to continue dining on the surrounding meal.
To precisely identify what these ion channels detect, Estrem and Flavell exposed worms to 20 different bacteria species known to be part of the worm's environment. They observed varying degrees of NSM activation across these bacteria. Through systematic chemical breakdown of bacterial components, the researchers progressively eliminated potential triggers, including DNA, lipids, proteins, and simple sugars.
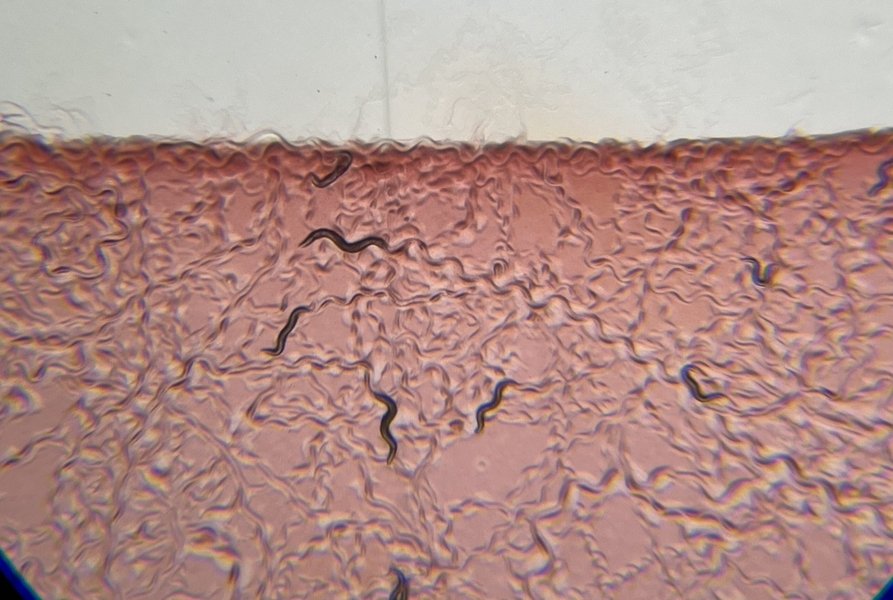

The breakthrough came when they identified that polysaccharide sugars coating many bacteria were responsible for driving NSM activation. Specifically, in gram-positive bacteria, a chemical called peptidoglycan activated NSM. For gram-negative bacteria, a different polysaccharide appeared to be the key trigger. The team demonstrated that these polysaccharides not only trigger NSM electrical activity but also promote the associated feeding and slowing behaviors. Crucially, genetically knocking out the ASICs abolished these responses, confirming that polysaccharide detection requires these specific ion channels.
Detecting Danger: The Prodigiosin Factor
Having identified what attracts worms to beneficial bacteria, the researchers turned their attention to danger signals. They focused on Serratia marcescens, a bacterium that can infect both worms and humans. Some strains of this bacterium produce a red pigment called prodigiosin, which makes them particularly lethal to worms.
The experiments revealed a sophisticated detection system: when NSM encountered non-pigmented bacteria, the neuron activated, and worms ingested the bacteria. However, when prodigiosin was present, NSM remained inactive, and worms avoided ingesting the dangerous bacteria. Adding prodigiosin to normally beneficial bacteria also suppressed NSM's usual response, indicating that the worms have evolved specific mechanisms to avoid this particular danger signal.
Implications for Broader Neuroscience
The molecular mechanisms identified in C. elegans may have broader relevance across species. "We developed a way of identifying these pathways by studying this organism that specializes in bacterial detection and displays robust responses," Flavell explains. "But there's no reason these pathways should be limited to C. elegans. The molecular players we identified are found in many species, including mammals."
This research provides a foundation for understanding how the gut microbiome communicates with the nervous system, potentially informing treatments for neurological conditions linked to bacterial imbalances. The specific identification of polysaccharides and prodigiosin as key signals opens avenues for targeted interventions that could modulate these neural pathways.
The study was supported by the National Institutes of Health, the McKnight Foundation, the Alfred P. Sloan Foundation, the Howard Hughes Medical Institute, and The Freedom Together Foundation. Beyond Estrem and Flavell, the paper's authors include Malvika Dua, Colby Fees, Greg Hoeprich, Matthew Au, Bruce Goode, and Lingyi Deng.
For those interested in exploring this research further, the Flavell Lab and the Picower Institute for Learning and Memory provide additional resources on this and related work. The full paper, "Identification of bacterial signals that modulate enteric sensory neurons to influence behavior in C. elegans," can be found in the journal Current Biology.

The findings represent a significant step toward understanding the complex dialogue between our nervous systems and the microbial communities that inhabit our bodies, potentially opening new avenues for therapeutic interventions targeting the microbiome-brain axis.
Comments
Please log in or register to join the discussion